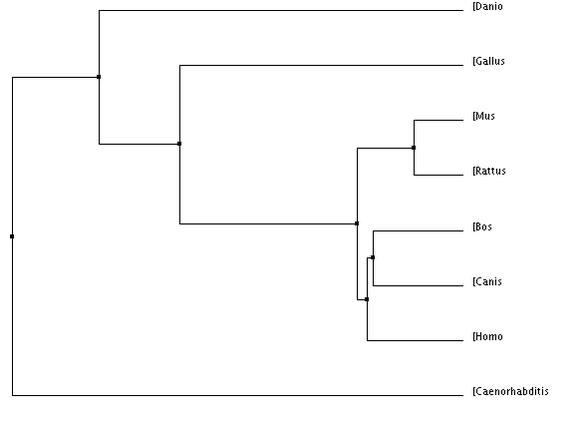
Phylogeny
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
Phylogeny is used to show the evolutionary relationship between organisms.
Constructing a Phylogenic Tree
Obtaining Protein Sequences. The first step in constructing a phylogenetic tree is obtaining the protein sequence of your gene of interest in its homologs. To use online tools, the protein sequence must be in FAFSA formatting.
|
|
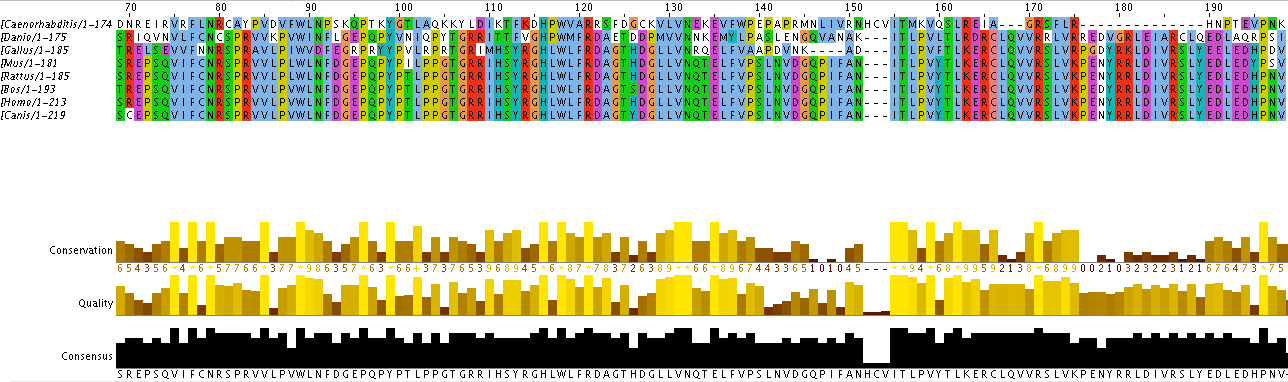
Sequence Alignment. With the protein's sequences of your homologs formatted, you can copy and paste it to an online tool called ClusterOmega. Cluster Omega will align your protein sequences and provide a document that can be copied and pasted into the program Jalview. Jalview can also be used to output your tree.
3. Interpeting Phylogenetic Trees.
|
To compare sequences
BLOSUM Martix Percent Identity |
Methods of drawing your tree
Neighbor Joining Average Distance |
Phylogeny of VHL
Conclusions |
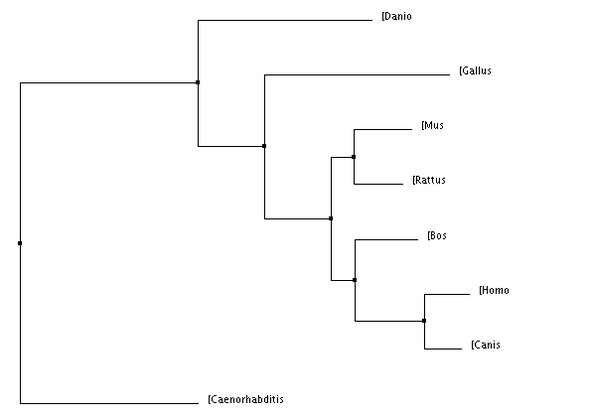
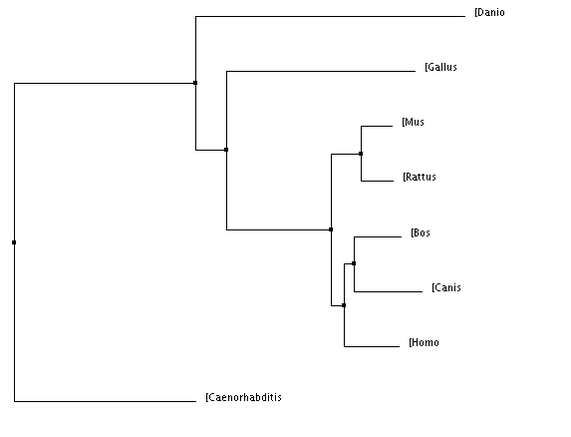
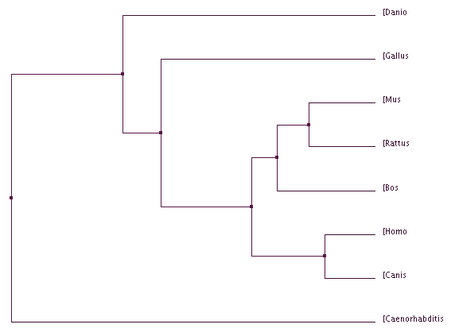
BLOSUM 64: scores the genes by the number of amino acids in common
Figure 1&2: The homo sapiens form of VHL is more closely related to dogs than other organisms. PIC (Percent Identity): scores the gene's sequence based on their similarity Figure 3&4: The homo sapiens form of VHL is more closely related to dogs and cows compared to other organisms. |